Project Status: Prevention-First Approach
So far, I'm fortunate to report zero varroa detections across all six hives. This system wasn't built in response to an infestation—it's a proactive, passion project designed to catch problems before they start. Think of it as an early warning system that complements my existing monitoring methods, particularly alcohol washes.
The goal was never just to detect varroa after it appears, but to build a continuous monitoring system that gives me peace of mind and catches the first signs of trouble weeks earlier than traditional methods alone.
The Problem: Varroa Destructor
Varroa mites are parasitic mites that attach to honeybees and feed on their hemolymph (bee blood). Left unchecked, they can devastate an entire colony within months. Early detection is absolutely critical—but manually inspecting thousands of bees is time-consuming and often catches infestations too late.
Traditional monitoring methods involve sticky boards, alcohol washes, or visual inspections during hive checks. These are invasive, labor-intensive, and only give you a snapshot of a single moment in time. We needed something better.
System Overview
System in Action
Here's what the system looks like in practice, from the mites we're detecting to the alerts in action.
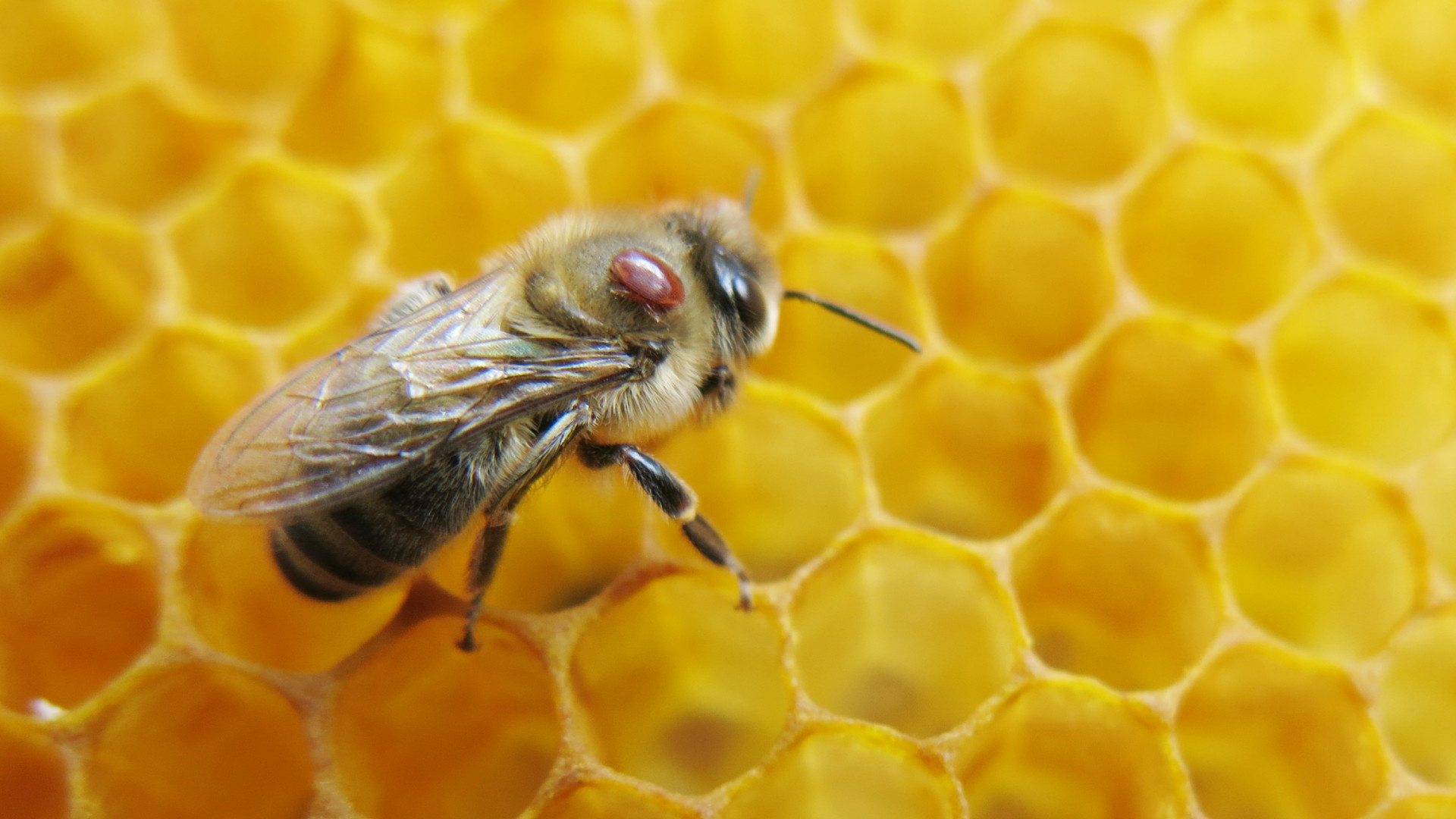

A varroa mite (reddish-brown) attached to a honeybee. These parasites are only 1-2mm but can devastate entire colonies.
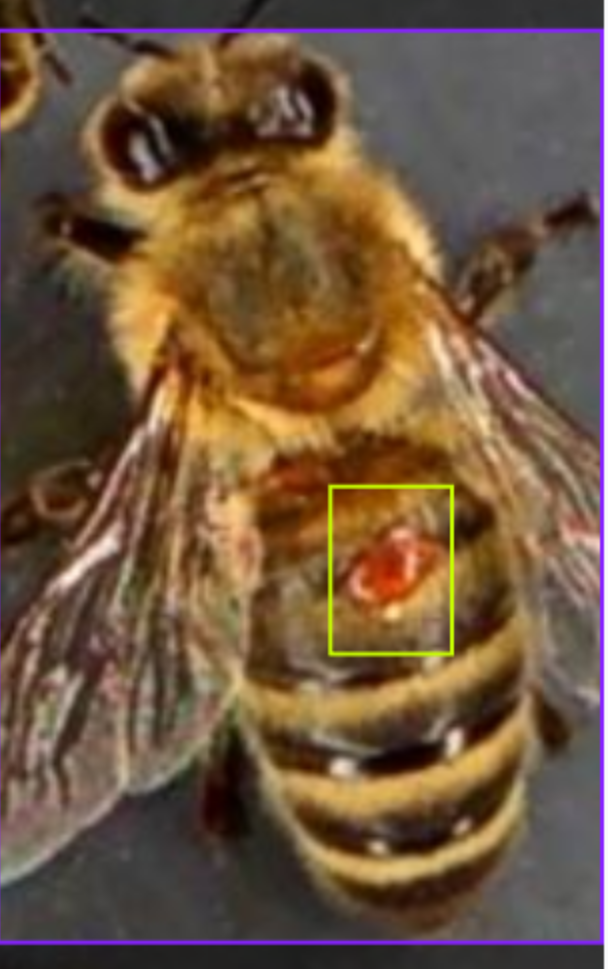

YOLOv11n model output showing detected mites with bounding boxes and confidence scores.
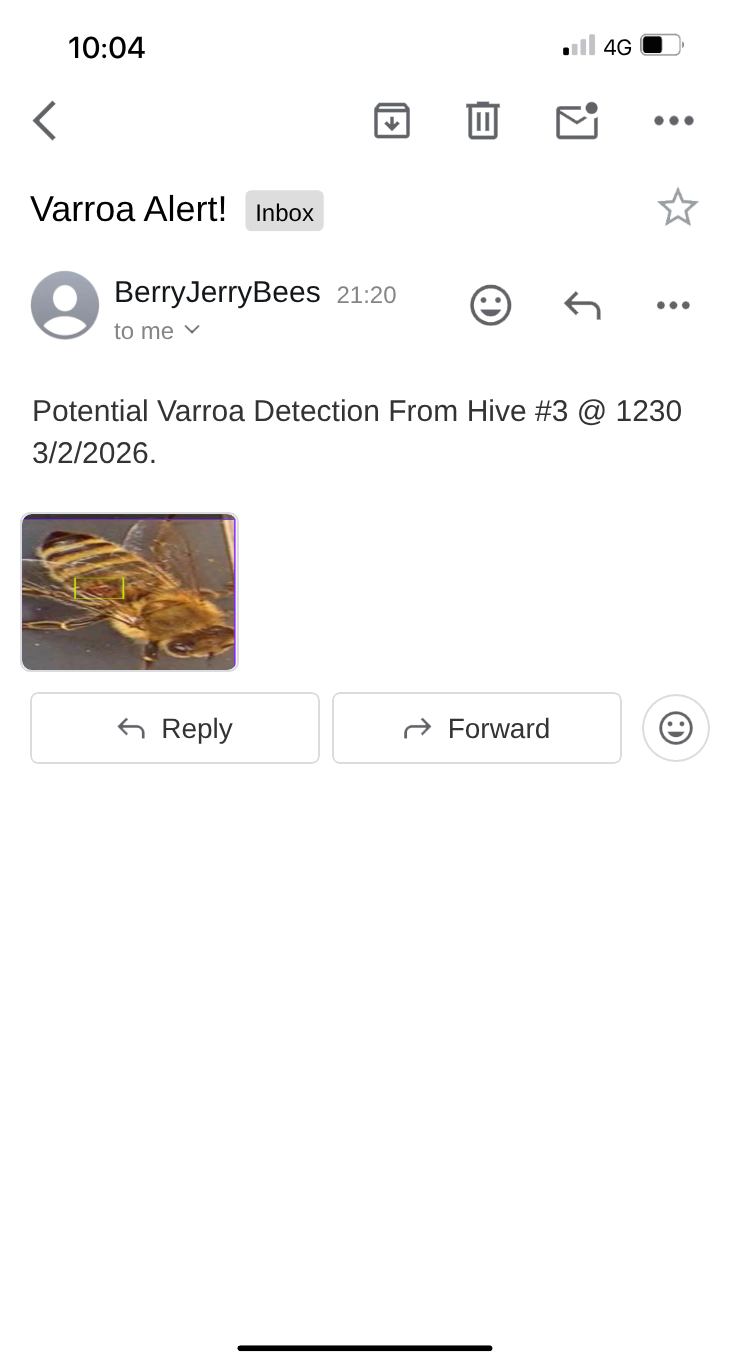
Automated alert received when the system detects potential varroa presence, with annotated image attached.
Hardware Setup
Each hive has its own monitoring station, completely solar-powered and weatherproof. Here's what goes into each setup:
- Camera: 4K resolution camera module (critical for catching tiny mites)
- Computer: Raspberry Pi 4 (handles image capture and uploads)
- Power: Solar panel + battery system (fully off-grid operation)
- Mounting: Custom 3D-printed housing positioned above hive entrance
- Storage: Local SD card backup + cloud upload to server
- Network: WiFi connection to home network for uploads
Camera Positioning
The cameras are mounted directly above the hive entrance, pointing down at a 45° angle. This captures bees as they enter and exit, giving us clear views of their backs—where varroa mites typically attach. The 4K resolution is essential because varroa mites are only 1-2mm in size.
Model Training
I used YOLOv11n (YOLO "nano" - the lightweight version) for object detection. Here's how I trained it:
1. Dataset Preparation
Found approximately 7,000 labeled images of varroa-infected bees on Ultralytics' open dataset. These images show bees with clearly visible varroa mites attached, along with bounding box annotations marking exactly where the mites are located.
Since I was working with two separate datasets (V1 and V2), I needed to merge them into a single master dataset. Here's the Python script I used to combine the validation sets while preventing filename conflicts:
Dataset Merging Script
import os
import shutil
from pathlib import Path
# Paths to source datasets
sources = [
Path(r'C:\Users\teapot\Documents\Projects\VarroaDetection\datasets\V1'),
Path(r'C:\Users\teapot\Documents\Projects\VarroaDetection\datasets\V2')
]
# Master dataset directories
master_val_img = Path(r'C:\...\master_dataset\val\images')
master_val_lbl = Path(r'C:\...\master_dataset\val\labels')
# Create master directories
master_val_img.mkdir(parents=True, exist_ok=True)
master_val_lbl.mkdir(parents=True, exist_ok=True)
# Common names for validation sets
val_names = ['val', 'valid', 'validation', 'test']
for src in sources:
found_in_src = False
for v_name in val_names:
img_dir = src / v_name / 'images'
lbl_dir = src / v_name / 'labels'
if img_dir.exists():
print(f"Found data in: {img_dir}")
found_in_src = True
for file in img_dir.iterdir():
if file.suffix.lower() in ['.jpg', '.jpeg', '.png']:
# Use source folder name as prefix to prevent conflicts
prefix = f"{src.name}_{v_name}_"
shutil.copy2(file, master_val_img / f"{prefix}{file.name}")
# Copy corresponding label file
label_file = lbl_dir / f"{file.stem}.txt"
if label_file.exists():
shutil.copy2(label_file, master_val_lbl / f"{prefix}{label_file.name}")
if not found_in_src:
print(f"!!! Warning: No validation folders found in {src}")
print(f"\nMerge complete. Total images: {len(list(master_val_img.glob('*')))}")
This script intelligently merges datasets by adding prefixes (like "V1_val_" or "V2_val_") to filenames, preventing any overwrites. It also handles different naming conventions for validation folders and ensures that both images and their corresponding label files are copied together.
2. Model Selection
Chose YOLOv11n because:
- Speed: Fast enough to process images in near real-time on a Raspberry Pi
- Accuracy: Good balance between detection accuracy and computational efficiency
- Size: Small model footprint suitable for edge deployment
3. Training Configuration
Training Parameters
Epochs: 100 Batch size: 16 Image size: 640x640 Augmentation: Enabled (flips, rotations, brightness adjustments) Validation split: 20% Hardware: NVIDIA GPU (local training)
Here's the complete training script I used. Running this on a laptop with an NVIDIA GPU took approximately 2-3 hours to complete all 100 epochs:
Training Script (train.py)
from ultralytics import YOLO
if __name__ == '__main__':
# Load the YOLOv11 nano model (optimized for speed)
model = YOLO('yolo11n.pt')
# Start training
model.train(
data=r'C:\Users\teapot\Documents\Projects\VarroaDetection\datasets\master_dataset\data.yaml',
epochs=100,
imgsz=640,
batch=16, # Adjusted for laptop GPU memory
device=0, # Use NVIDIA GPU (0 = first GPU)
workers=4, # Data loading threads
name='varroa_yolo11n_100epochs'
)
Key parameters explained:
- batch=16: Number of images processed simultaneously. Lower values use less GPU memory but train slower
- device=0: Tells YOLO to use the first NVIDIA GPU. Use 'cpu' if no GPU available
- workers=4: Number of parallel threads for loading images. Keep low on Windows to avoid overhead
- imgsz=640: All images resized to 640×640 pixels during training for consistency
Training Tips
GPU Memory Issues? If you get "CUDA out of memory" errors, reduce the batch size to 8 or even 4. Training will take longer but won't crash.
No GPU? Set device='cpu' in the training script. It'll be much slower (10-20x),
but it works. Consider using Google Colab's free GPU for faster training.
Monitoring Training: YOLO automatically saves training curves, metrics, and model checkpoints
to runs/detect/varroa_yolo11n_100epochs/ after each epoch. Check these to see how your model is improving!
4. Results
After training, the model achieved:
- Precision: ~94% (when it says "varroa detected," it's right 94% of the time)
- Recall: ~91% (catches about 91% of actual varroa infestations)
- Inference time: ~200ms per image on Raspberry Pi
Model Performance Metrics
After 100 epochs of training, here's how the YOLOv11n model performed. These metrics help us understand not just whether the model works, but how well it works and where it might need improvement.
Training Results Overview
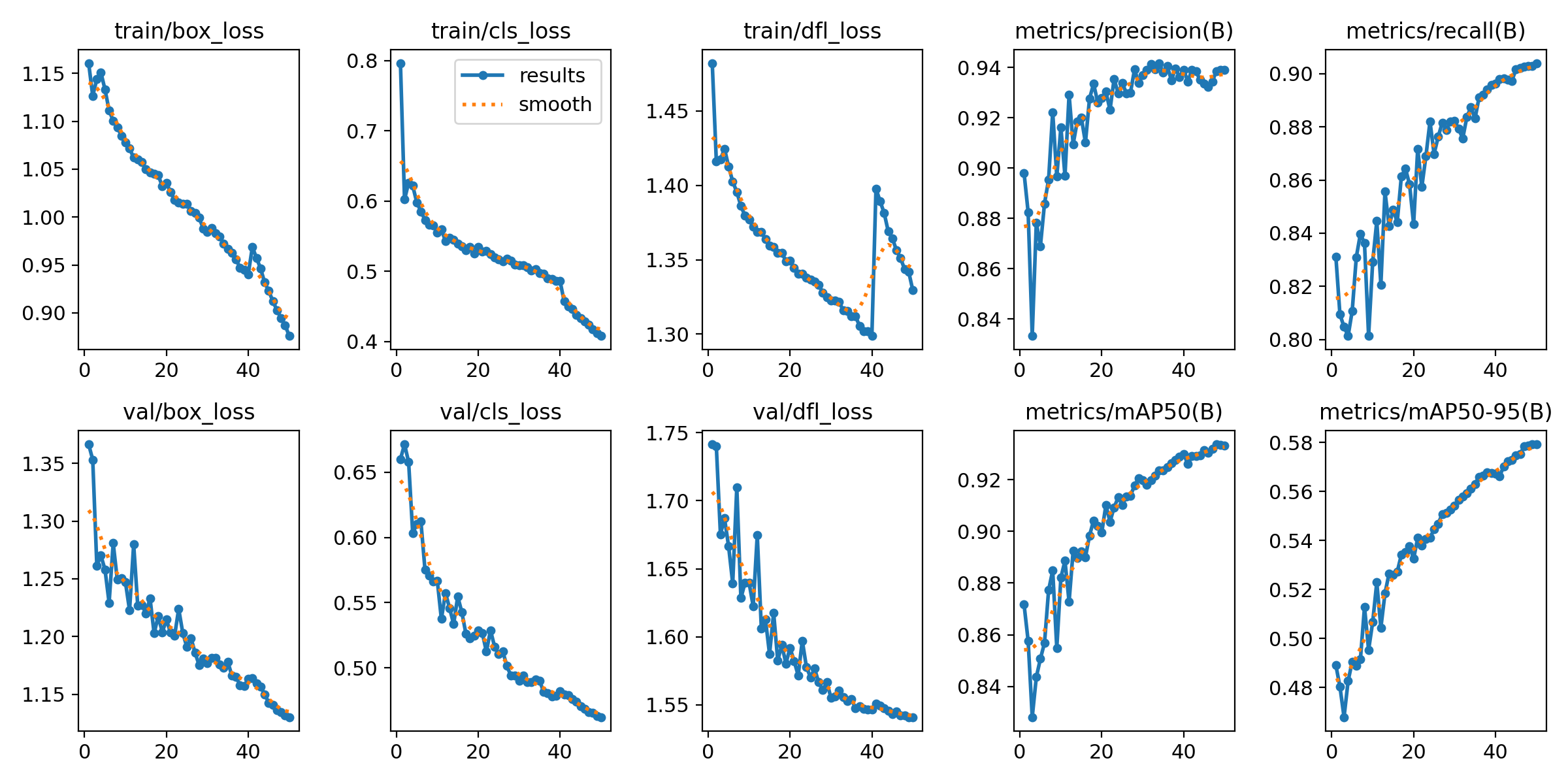
Loss curves and performance metrics across all 100 training epochs. Notice how the model steadily improves and stabilizes.
The results chart shows multiple key metrics tracked during training. The top row shows three types of loss (box, class, and DFL) decreasing over time—this means the model is learning. The bottom row shows validation losses, which tell us the model isn't just memorizing the training data but can generalize to new images.
Understanding the Metrics
The right side of the training results shows the most important numbers:
- Precision (94%): When the model says "I found varroa," it's correct 94% of the time
- Recall (91%): The model catches about 91% of all varroa mites present in images
- mAP50 (93%): Overall detection accuracy at 50% confidence threshold
- mAP50-95 (58%): Stricter accuracy metric across multiple thresholds
What This Means in Practice
With 94% precision and 91% recall, this model strikes a good balance. High precision means I won't get flooded with false alarms about varroa that isn't there. High recall means the system won't miss many real infestations. For a beekeeping application where early detection is critical, this is exactly the balance we want.
Precision-Confidence Relationship
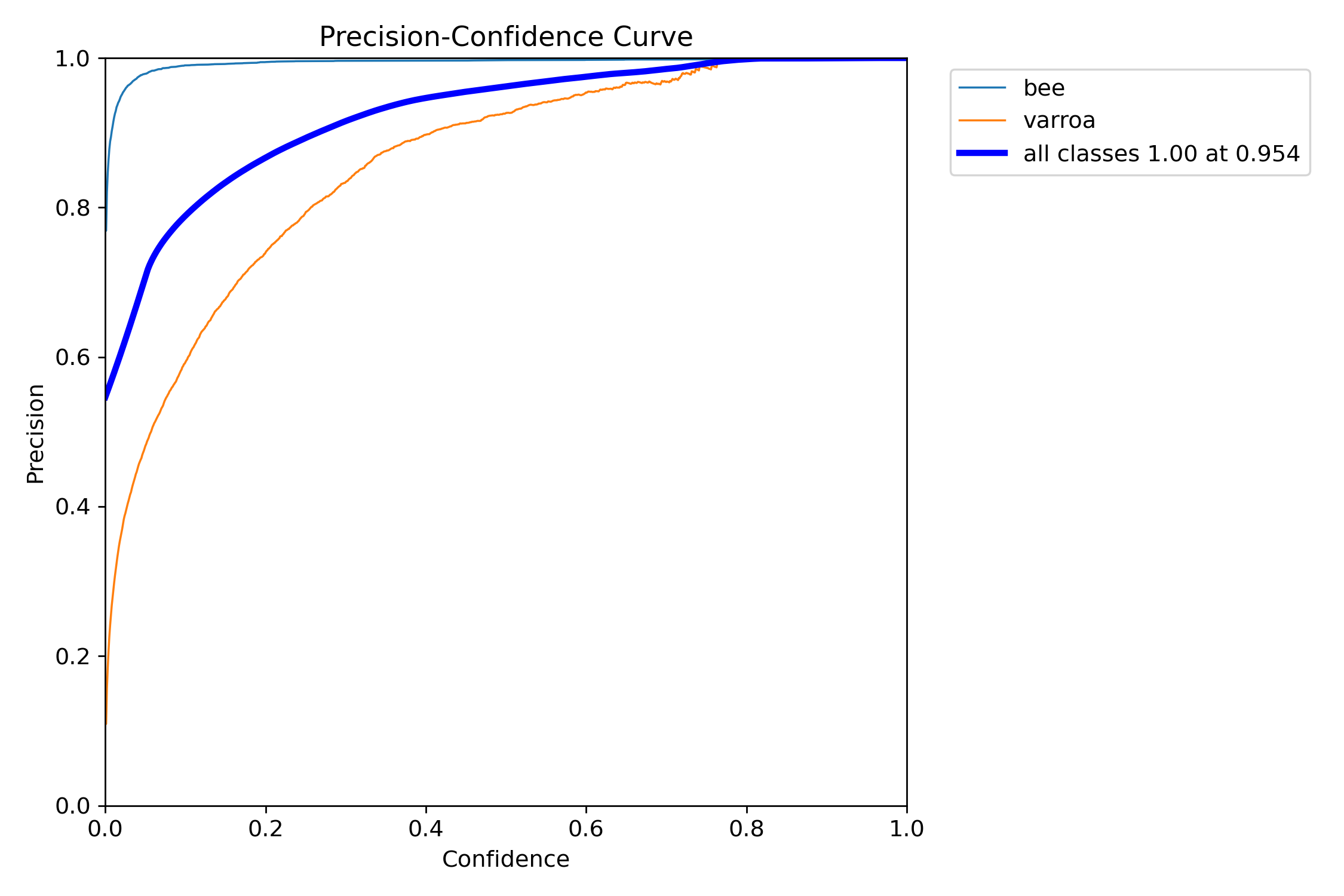
How precision changes as we adjust the confidence threshold. Higher thresholds mean fewer false positives.
This curve is crucial for tuning the system. It shows that the "bee" class (blue) achieves near-perfect precision very quickly, while "varroa" (orange) requires moderate confidence (around 0.4-0.5) to reach peak precision. The steep rise in the varroa curve means the model quickly becomes confident when it detects a mite—exactly what we want.
Precision-Recall Trade-off
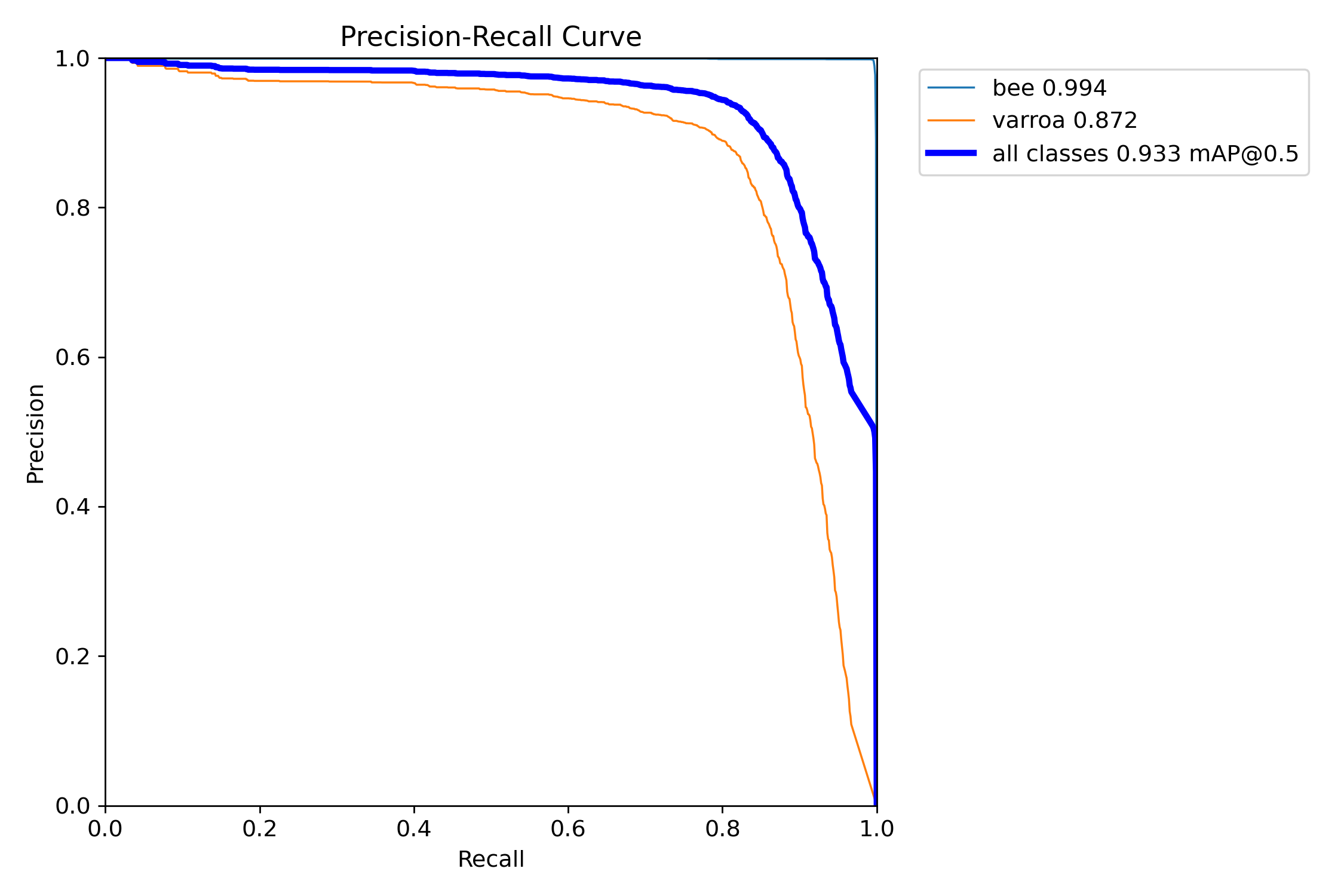
The classic ML trade-off: high precision with good recall. [email protected] of 93.3% is excellent for this application.
This is the classic machine learning trade-off curve. For varroa detection, we achieved an mAP of 87.2%, meaning the model maintains high precision across a wide range of recall values. The "all classes" curve (blue) shows an overall [email protected] of 93.3%—a strong result indicating the model generalizes well.
Recall-Confidence Analysis
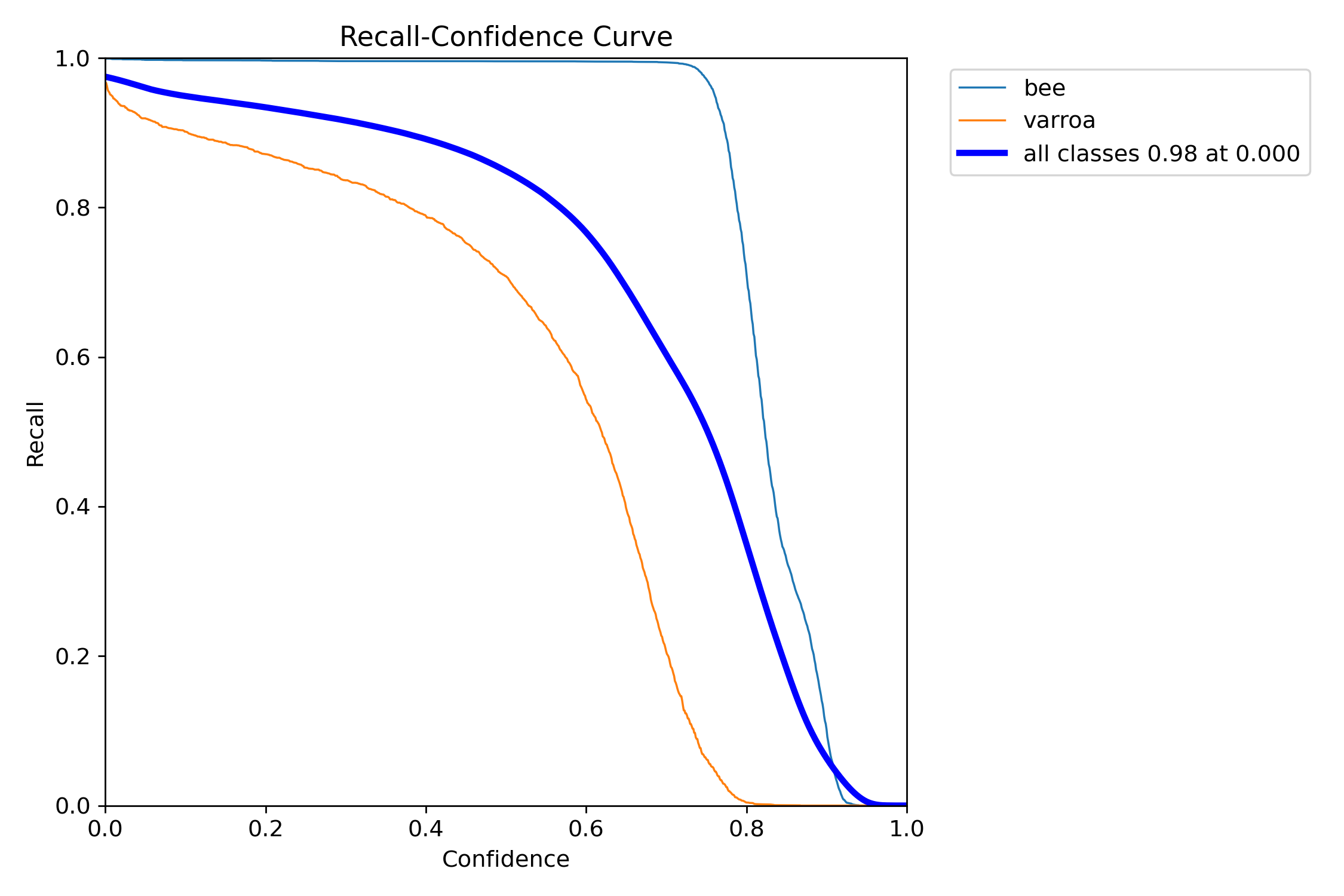
At very low confidence thresholds, we catch almost everything—but at what cost in false positives?
This curve shows how many varroa mites we catch at different confidence thresholds. At near-zero confidence, we catch 92% of all mites (high recall). But as we increase the threshold to reduce false positives, recall drops. The sweet spot for this system is around 0.5-0.7 confidence, where we still catch most mites while filtering out obvious false detections.
Confusion Matrix Analysis
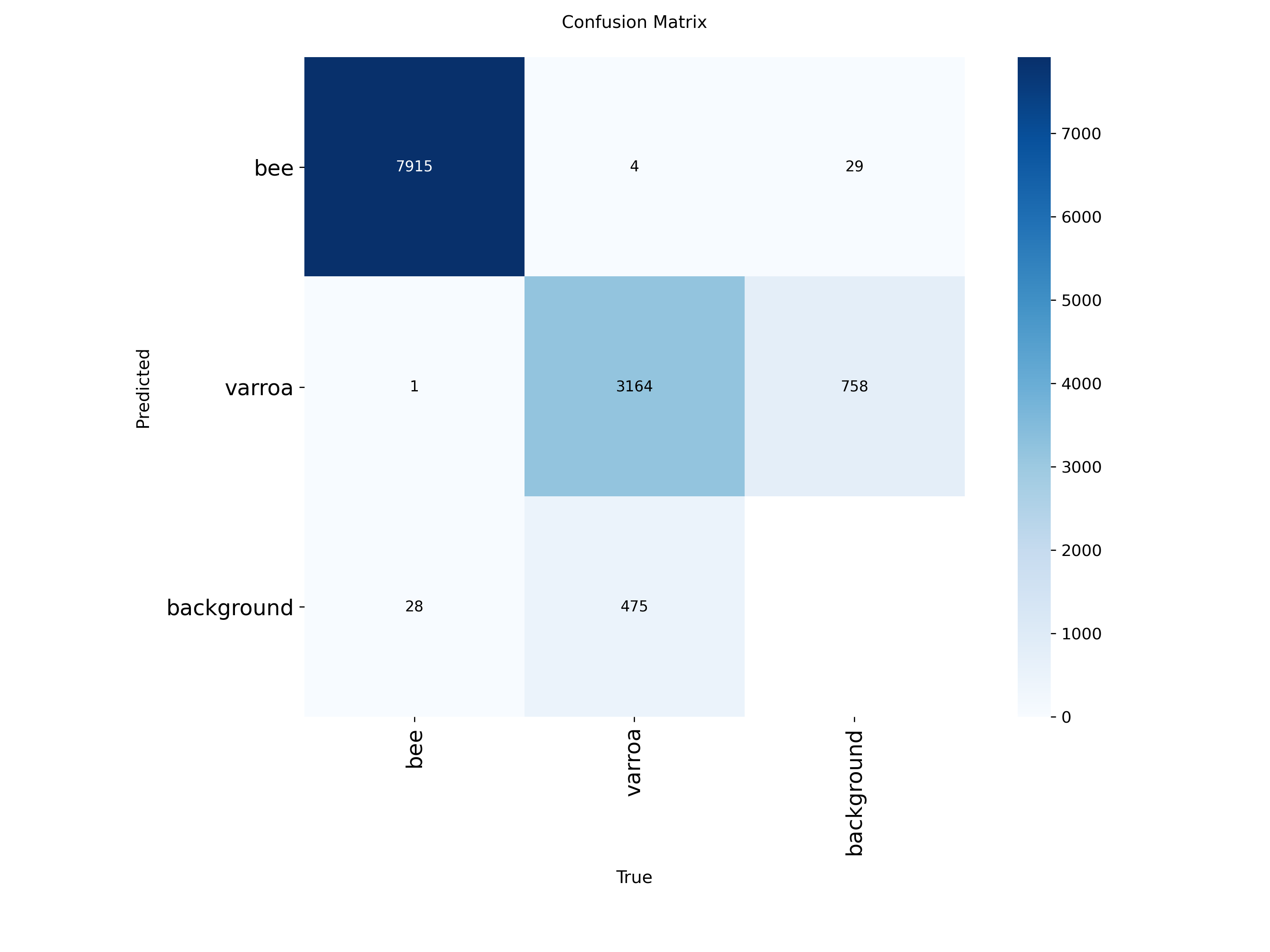
Where the model succeeds and where it struggles. Most errors are varroa mislabeled as background.
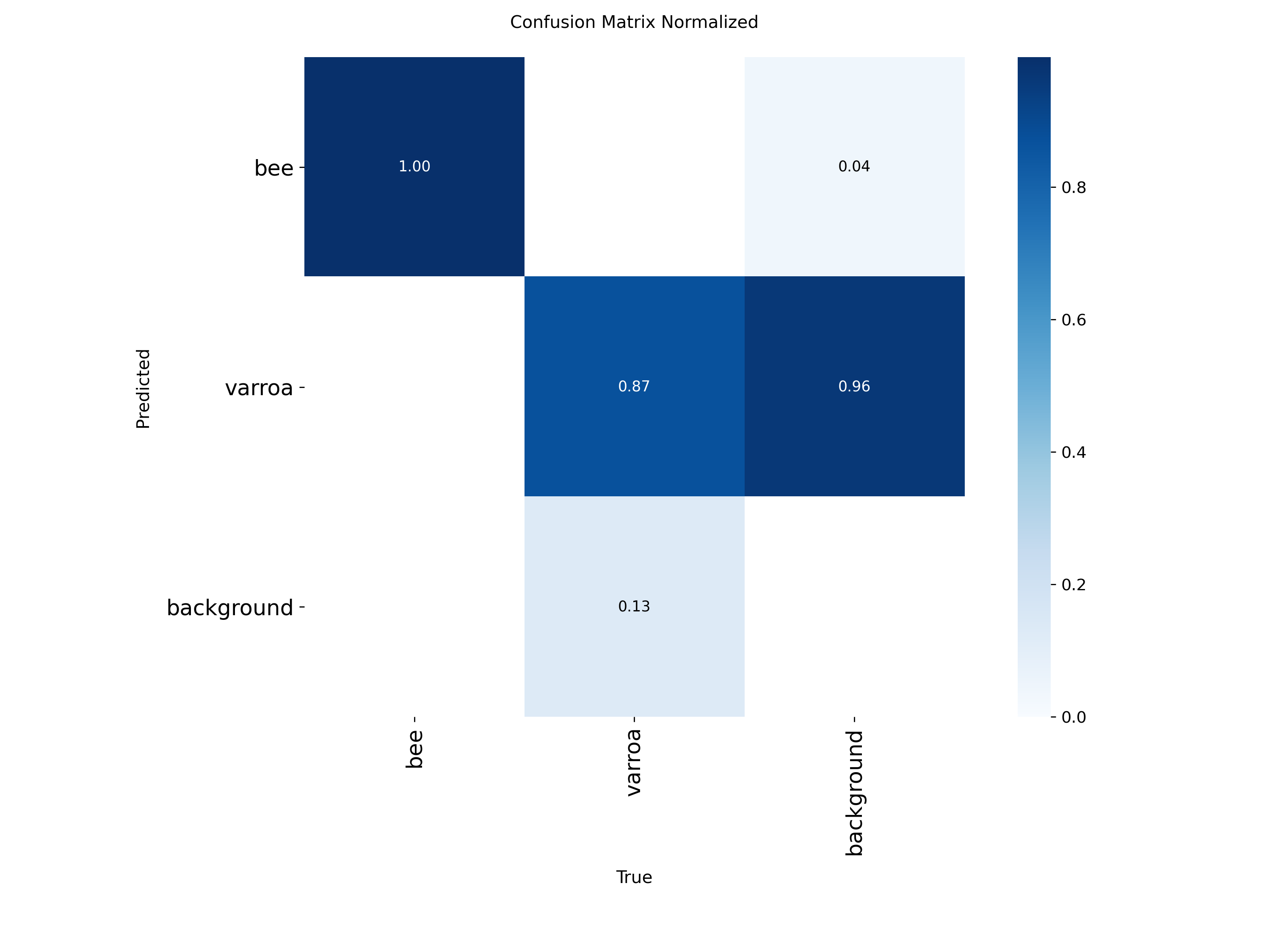
Proportional view showing that 87% of varroa instances are correctly identified, with most errors being false negatives.
The confusion matrices reveal the model's strengths and weaknesses:
- Bees (100% accuracy): The model almost never misidentifies bees—7,915 correct out of 7,948 instances
- Varroa (87% recall): Of 3,923 varroa instances, the model correctly found 3,164. The 758 false negatives (missed detections) are mostly mites misclassified as "background"
- Background class: Some background is mistaken for varroa (475 instances), which contributes to false positives
The normalized matrix shows that 96% of varroa predictions are classified as varroa or background, with very few confused as bees. This is ideal because it means the primary source of error is missing mites (false negatives) rather than hallucinating them (false positives). For a monitoring system, it's better to miss a few mites and catch them later than to constantly trigger false alarms.
Detection Distribution & Label Quality
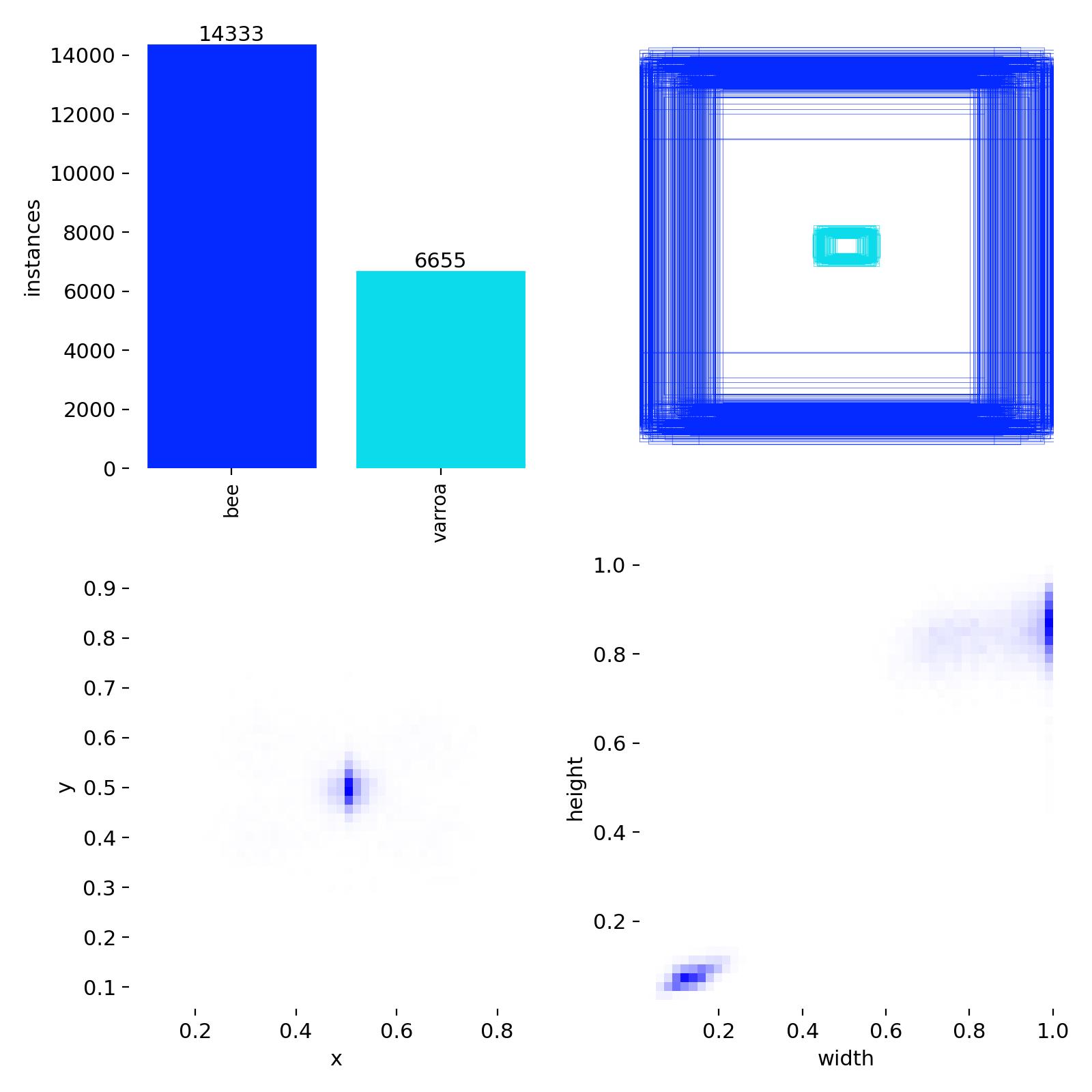
Class balance (14,333 bees vs. 6,655 varroa) and bounding box size/position distributions from the training set.
This diagnostic chart reveals important characteristics of the training dataset:
- Class imbalance: About 2:1 ratio of bees to varroa, which is realistic for detection scenarios
- Bounding box positions: Varroa mites cluster around x=0.5, y=0.5 (center of images), while bees are more distributed
- Box dimensions: Varroa bounding boxes are concentrated in the small size range (width/height ~0.05-0.15), confirming these are tiny objects
The small, centered distribution of varroa bounding boxes explains why high-resolution 4K cameras are essential— these mites occupy only a tiny fraction of each image frame.
What These Metrics Mean for Deployment
Based on these results, I've configured the production system with a confidence threshold of 0.70. This means the model must be at least 70% confident before triggering an alert. At this threshold:
- False positives are minimized (I won't get alerts for every bit of pollen or shadow)
- True positives remain high (we catch the vast majority of actual varroa)
- The system leans conservative—better to check a hive based on a real alert than ignore a genuine infestation
The confusion matrix confirms this is the right approach: most errors are missed detections (which get caught in the next photo 30 minutes later) rather than false alarms (which would erode trust in the system).
System Architecture
Here's how the entire system works, from camera to inbox:
graph TD
A[4K Camera] -->|Every 30 min| B[Raspberry Pi]
B -->|Capture Image| C[Local Storage]
C -->|Upload via WiFi| D[Server Directory]
D -->|Cron Job Every 30min| E[Analysis Script]
E -->|Load Image| F[YOLOv11n Model]
F -->|Run Inference| G[Varroa Detected?]
G -->|No| H[Log and Continue]
G -->|Yes| I[Generate Alert]
I -->|Email| J[Beekeeper Inbox]
I -->|Include| K[Annotated Image]
I -->|Include| L[Hive Identifier]
I -->|Include| M[Detection Confidence]
J -->|Manual Review| N[Inspect Hive]
Detailed Workflow
Step 1: Image Capture (Every 30 Minutes)
A cron job on each Raspberry Pi triggers the camera every 30 minutes. The Pi captures a 4K image of the hive entrance, stamps it with a timestamp and hive ID, and saves it locally.
Crontab Entry (Raspberry Pi)
*/30 * * * * /home/pi/capture_image.sh
Step 2: Upload to Server
Each Raspberry Pi then uploads the image to a central server directory via rsync or SFTP. The filename includes the hive name and timestamp for easy identification.
Example Filename
padme_2026-02-08_0830.jpg galadriel_2026-02-08_0830.jpg frigga_2026-02-08_0830.jpg
Step 3: Automated Analysis
A separate cron job on the server runs every 30 minutes, scanning the upload directory for new images. For each new image:
- Load the image into memory
- Pass it through the YOLOv11n model
- Parse detection results (bounding boxes, confidence scores)
- If varroa mites are detected above a confidence threshold (e.g., 70%), flag the image
Pseudocode: Analysis Script
for each new_image in upload_directory:
results = model.predict(new_image)
if results.detections.confidence > 0.70:
annotated_image = draw_boxes(new_image, results)
hive_name = extract_hive_from_filename(new_image)
timestamp = extract_timestamp(new_image)
send_alert_email(
to="[email protected]",
subject=f" Varroa Alert: [hive_name]",
body=f"Varroa detected at [timestamp]",
attachment=annotated_image
)
log_detection(hive_name, timestamp, confidence) Step 4: Email Alert
When varroa is detected, I receive an email with:
- Subject line: " Varroa Alert: [Hive Name]"
- Body: Timestamp, confidence score, and detection details
- Attachment: The original image with bounding boxes drawn around detected mites
- Hive location: Which queen's hive needs attention
Why Email Alerts?
Email is simple, reliable, and doesn't require maintaining a separate dashboard or app. I get push notifications on my phone, can view the image immediately, and have a searchable history of all detections. Plus, I can forward alerts to other beekeepers if I'm away.
Step 5: Manual Verification & Action
Once I receive an alert, I:
- Review the image: Check if the detection is a true positive (sometimes pollen or debris can trigger false positives)
- Inspect the hive: Do a physical inspection within 24 hours
- Take action: If confirmed, treat the hive immediately (oxalic acid vaporization, formic acid strips, etc.)
- Update records: Log the detection and treatment in my hive management system
Data Flow
Here's a simplified view of how data moves through the system:
sequenceDiagram
participant C as Camera
participant RPi as Raspberry Pi
participant S as Server
participant M as ML Model
participant B as Beekeeper
C->>RPi: Capture image (every 30min)
RPi->>RPi: Save locally
RPi->>S: Upload via WiFi
S->>S: New images detected
S->>M: Run inference
M->>M: Analyze for varroa
alt Varroa detected
M->>S: Return detections
S->>B: Send email alert
S->>S: Log event
else No varroa
M->>S: No detections
S->>S: Log clear result
end
Real-World Impact & Results
What's Working Well
- Continuous peace of mind: 24/7 monitoring without disturbing the hives
- Complements alcohol washes: Visual monitoring between invasive sampling
- Non-invasive: No need to open hives or stress bees for routine checks
- Historical data: Building a baseline dataset of healthy hive entrance activity
- Scalable: Easy to add more cameras as the apiary grows
- Learning experience: Deep dive into computer vision and edge computing
Challenges & Ongoing Improvements
- False positives: Pollen, wood debris, and bee shadows can trigger alerts
- Weather dependency: Heavy rain or fog can obscure the camera
- Night detection: Currently daylight-only; considering IR LEDs for night vision
- Model refinement: Need more real-world varroa images from my specific setup
- Power management: Winter months with less sunlight can be challenging
Next-Generation Improvements
K-Means Color Clustering for False Positive Reduction
One of the biggest challenges with the current system is false positives—pollen, shadows, and debris can sometimes be mistaken for varroa mites. The next major improvement I'm working on is using K-means clustering to analyze the color profile of detected objects.
How K-Means Will Help:
- Color signature analysis: Varroa mites have a distinct reddish-brown color (~RGB 139, 69, 19)
- Cluster dominant colors: Extract the main colors from each detected bounding box
- Compare against varroa profile: Check if the dominant cluster matches known varroa coloration
- Secondary validation: Only trigger alerts if both YOLO detection AND color clustering agree
- Reduce false positives: Filter out yellow pollen, dark shadows, and wood fragments
Conceptual Workflow
# After YOLO detects potential varroa
if yolo_detection.confidence > 0.70:
# Extract the detected region
cropped_region = image[bbox.y1:bbox.y2, bbox.x1:bbox.x2]
# Apply K-means clustering (k=3 to find dominant colors)
kmeans = KMeans(n_clusters=3)
colors = kmeans.fit(cropped_region.reshape(-1, 3))
dominant_color = colors.cluster_centers_[0]
# Check if dominant color matches varroa profile
varroa_color_range = ([120, 50, 10], [160, 90, 40]) # RGB range
if is_color_in_range(dominant_color, varroa_color_range):
confidence_boost = 1.2 # Increase confidence
send_alert()
else:
# Likely false positive - log but don't alert
log_potential_false_positive()
This approach should dramatically reduce false positives from yellow pollen (which clusters toward RGB ~255, 200, 0) and dark shadows (which cluster toward low RGB values). By combining computer vision object detection with color analysis, we get a much more robust detection system.
Other Planned Enhancements
- Multi-class detection: Expand model to detect other pests (small hive beetle, wax moths)
- Bee counting: Track hive population and foraging activity over time
- Temperature sensors: Integrate thermal monitoring for brood health
- Edge processing: Run inference directly on Raspberry Pi to reduce server load
- Historical trend analysis: Detect patterns that precede infestations
- Mobile app: Real-time push notifications instead of email
- Other Detection Models: Explore DETR or YOLO v12 to fine tune and minimise false positives
Technology Stack
graph LR
A[Hardware Layer] --> B[Raspberry Pi 4]
A --> C[4K Camera Module]
A --> D[Solar Panel System]
E[Software Layer] --> F[Python 3.11]
E --> G[Ultralytics YOLOv11]
E --> H[OpenCV]
E --> I[PIL/Pillow]
J[Infrastructure] --> K[Linux Server]
J --> L[Cron Jobs]
J --> M[SMTP Email]
J --> N[rsync/SFTP]
O[ML Pipeline] --> P[Model: YOLOv11n]
O --> Q[Dataset: ~7000 images]
O --> R[Framework: PyTorch]
Questions? Want to Build Your Own?
I'm happy to share more details about the setup, code, or answer questions from fellow beekeepers interested in implementing similar systems. This kind of technology could be game-changing for small-scale beekeepers who don't have time for constant manual monitoring.
Get in touch: [email protected]